Structural Bioinformatics of Proteins, taught by Senior Lecturer in Biology Craig Smith, allows undergraduate students the opportunity to use cutting-edge bioinformatics tools to evaluate protein structures. It also exposes them to the process of being a scientist.
A refreshing swim in a lake on a warm day may be tempting, but you might want to think twice before going swimming this summer. Lurking in warm freshwater are brain-eating amoebae capable of doing what their name implies – they eat brain.
The good news is your chances of falling victim to Naegleria fowleri are very small. The bad news is the majority of infections are fatal. There is no effective treatment to reverse the damage the amoeba inflicts when it travels up the nasal cavity and burrows its way to the frontal lobe.
Now a collaboration between structural biologists at the University of Washington in Seattle and undergraduate students at Washington University in St. Louis has generated data published in PLoS One in March that will facilitate drug discovery for this neglected disease.
“My students are providing a service. They are performing analyses that can be accessed and leveraged by scientists to help design drugs or small molecules that will target a protein on the amoeba, as is the case in this paper. They are contributing to the drug discovery process,” said Craig Smith, senior lecturer in biology, who teaches students enrolled in Structural Bioinformatics of Proteins (Bio4525) to use cutting-edge bioinformatics tools to evaluate protein structures.
Scientific training
In 2016, the Seattle Structural Genomics Center of Infectious Disease (SSGCID) consortium, which creates protein structures for the scientific community, was looking for an educational partner. Smith was looking for a research partner. And a collaboration was born; Smith designed a course where students perform bioinformatics analyses using protein structures.

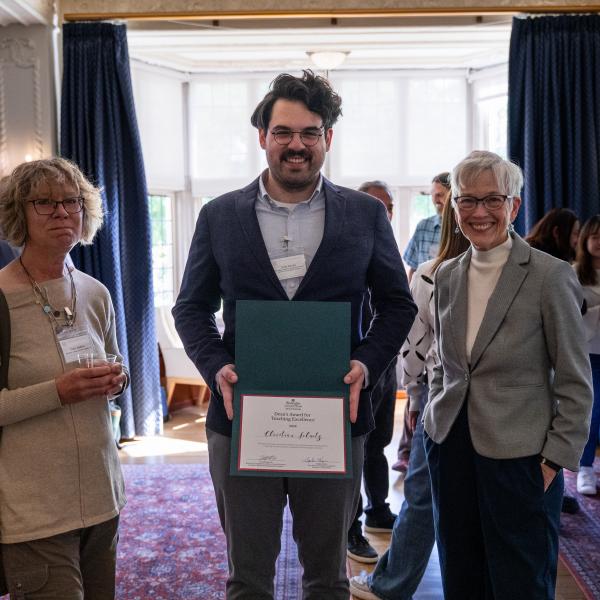
Jenna Goldstein, Ryan Young, Craig Smith
The students perform scientific inquiry to identify what we already know about the protein in the scientific literature. That inquiry helps them define the questions they want to answer.
“When students arrive at WashU, the biggest transition they make from high school, at least in STEM, is to think about open-ended questions that don’t have definitive answers. Many of them are thinking about these big questions and wrestling a big project down to its smaller parts for the first time,” said Smith.
The students take ownership of their projects – each one conducting original work and making a significant contribution to the scientific community. In the process, they learn how to critically evaluate scientific information, write clearly and present their ideas in group meetings. They quickly learn how to be independent researchers and thinkers.
They also learn to write a scientific manuscript. But Smith makes it clear that publishing a high-impact paper is not the primary goal.
“The goal for the students is not to publish in Nature, Science or Cell but to learn about the process of writing a publication-quality manuscript,” said Smith, who has fond memories of publishing his first scientific paper as an undergraduate student. Publishing in any journal is a proud moment for these students and, for many, their first foray into conducting independent scientific research.
Bio4525 students to the rescue
Three years into the course, Smith met Kayleigh Barrett, a researcher in the department of medicine, division of allergy and infectious diseases at the University of Washington and lead author on the PLoS One publication, during an update meeting with the SSGCID consortium. Impressed by the students’ work, Barrett enlisted their help to analyze protein structures from pathogens that cause neglected disease, focusing on brain-eating amoebae.
But Smith had a dilemma. It was now spring of 2019, and the class was over.
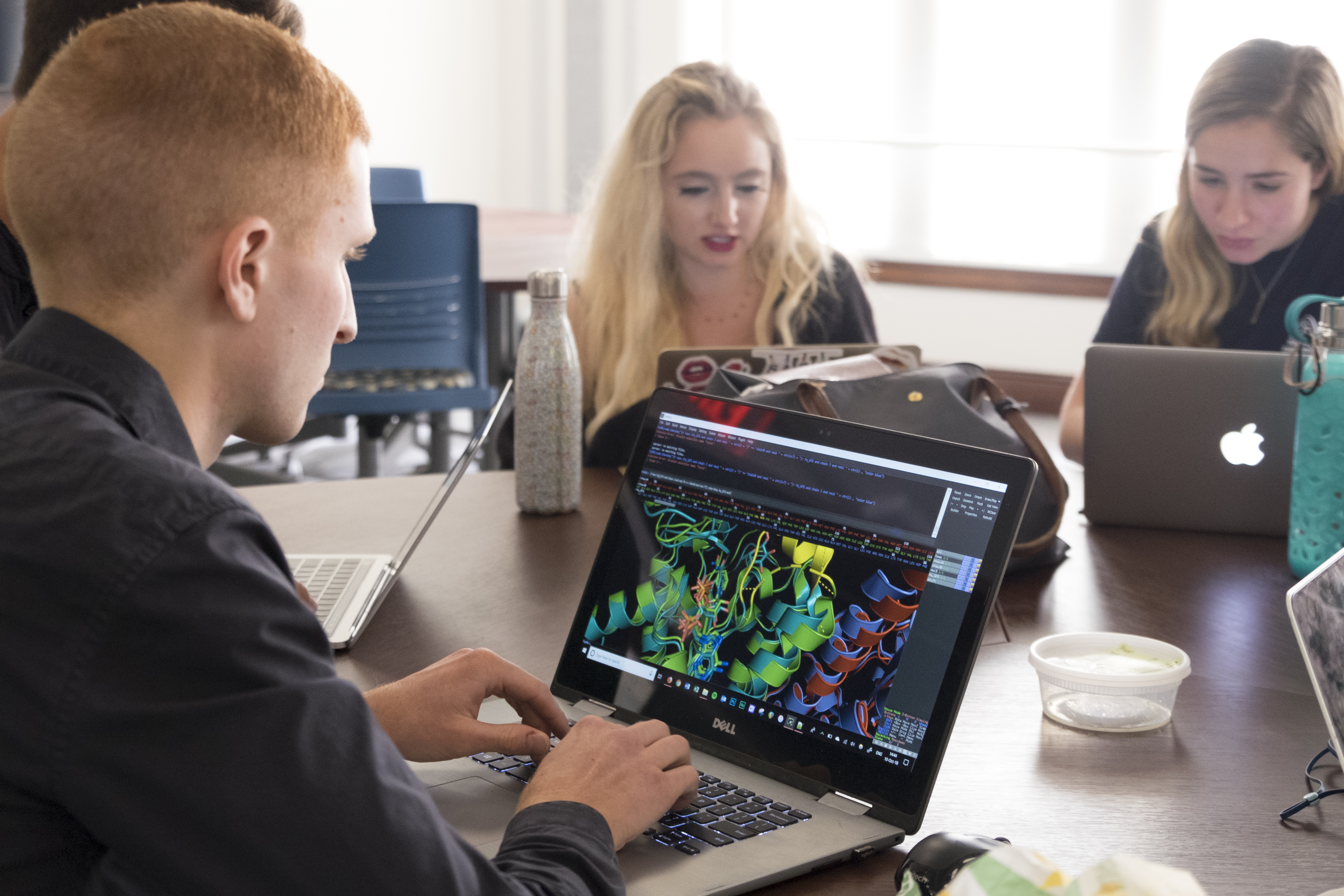

in 2018. Left to right starting at the front: Bram Osterhout, Caroline
Francis, and Becca Marks
Smith offered the project to the students in his Fall 2018 course. But this time, the students were on their own and expected to apply what they learned in the fall to perform the analysis. The response was overwhelming; the students were eager to independently execute the research in their free time.
They performed structural alignments with human versions of five Naegleria fowleri protein structures to identify differences. “Those differences are critical for developing a good drug target that affects the brain-eating microbe without causing side effects in humans,” explained Smith.
The students put together the figures and wrote up their findings for the paper. Six of Smith’s former students are authors on the paper, including Jenna Goldstein, Jared Lassner, Bram Osterhout, Nathan Tran, Lily Xu, and Ryan Young. They participated in Bio4525 in 2018 as juniors or seniors.
“This paper is a tribute to my students who over a semester developed into independent thinkers, scientists,” said Smith.
And proof that a side project waiting to be analyzed in a lab somewhere, anywhere might be the perfect collaboration for Smith and the next generation of Bio4525 students.